|
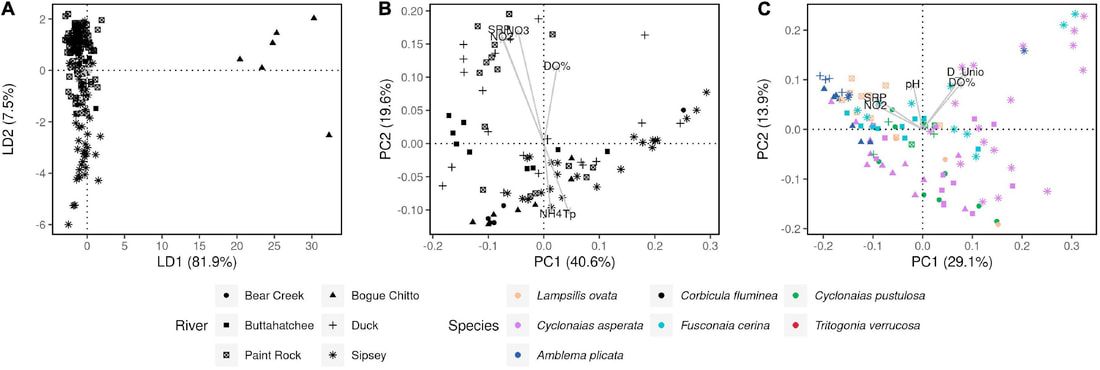
We just published a paper integrating population genomics (RADtag data generated by PhD student Jamie Bucholz) with microbiome data (from the Ole Miss DoB team) from Corbicula fluminea populations across the Mobile and TN river basins. This critter has some intriguing mechanisms of inheritance creating some unusual population genetic data, but overall we were able to pull out some interesting genetic patterns of structure in the species among rivers. The microbiome however, was unrelated to genetic structure of the clams, but was also structured by space and environment, and there was also seemingly an effect of the native mussel population on the C. flumina microbiome. The data suggest the microbiome may assist Corbicula in resource acquisition and facilitate invasion, but the processes structuring microbes and genetics of the populations differ during the invasion process.
Chiarello, M., Bucholz, J. R., McCauley, M., Vaughn, S. N., Hopper, G. W., Sánchez González, I., Atkinson, C.L., Lozier, J. D., Jackson, C. R. (2022). Environment and Co-occurring Native Mussel Species, but Not Host Genetics, Impact the Microbiome of a Freshwater Invasive Species (Corbicula fluminea) . Frontiers in Microbiology. https://www.frontiersin.org/article/10.3389/fmicb.2022.800061 Comments are closed.
|
Lozier Lab NewsDispatches from the lab and field! Archives
March 2023
Categories
All
|
 RSS Feed
RSS Feed